About
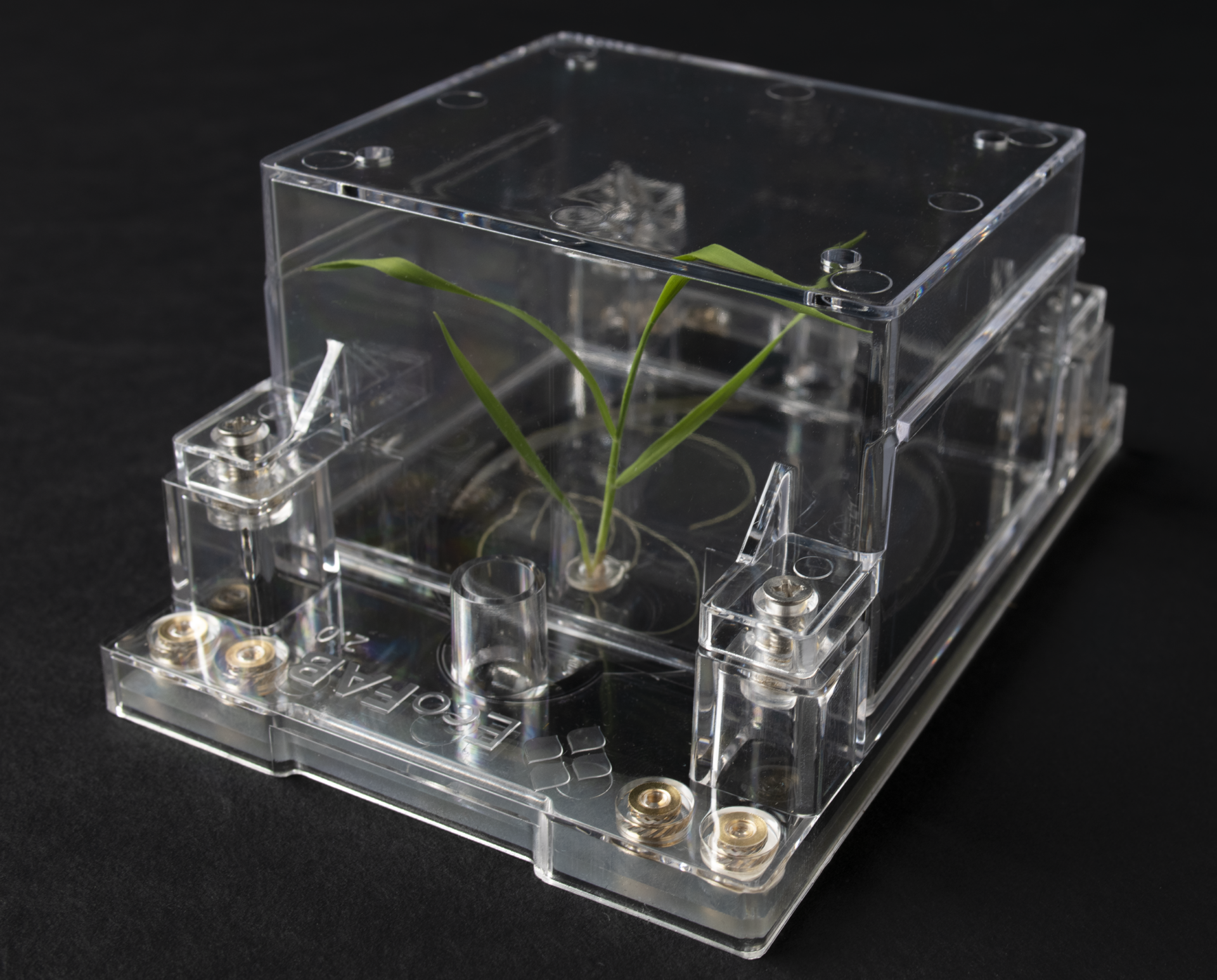
Fabricated ecosystems (EcoFABs) consist of an autoclavable physical chamber (enabling gnotobiotic studies), a target host organism (optional), growth medium (e.g. soil, sand, media, etc.), and the microbial communities to be tested. Great effort is made to develop standardized and reproducible systems to enable scientists to build on each other’s work. The exact format of the EcoFAB depends on the experimental system and scientific focus, but typically includes capabilities for imaging and omics analysis.
The initial design of EcoFAB 1.0 relied on 3D printing of molds used to cast Polydimethylsiloxane (PDMS). The greatly improved EcoFAB 2.0 design is created with standardized mass production by injection molding and thus does not require microfabrication by users.
Current EcoFAB Projects
Explore ongoing initiatives involving the EcoFAB devices.
TEAMS m-CAFEs
The EcoBOT
Videography: Marilyn Sargent
EcoFAB Fabrication and Development
EcoFAB is an initiative that aims at building model laboratory ecosystems across worldwide research laboratories to study environmental microbiomes for investigation of soil carbon cycling and low input sustainable crop production. Through this initiative, a lab scale device or fabricated ecosystem has been developed to study soil-plant-microbe interactions in a reproducible and controllable system.
Fabricated ecosystems, or EcoFABs, hold promise for replicating important ecosystem dynamics while simultaneously streamlining studies by defining testable and reproducible parameters. Used with microbiomes, scientists will be able to define principles for community assembly and structure, understand the functions of genes, microbes, and metabolomes, and predict microbiome health and trajectory. EcoFABs have the potential to bring together communities of scientists working on shared systems, enabling more effective knowledge transfer and the ability to build upon previous findings. By expanding our understanding of the assembly, structure, and functions of microbiomes, significant scientific and technical advances will be made in studies of biomedical technologies, environmental health, agriculture, energy, and nutrient cycling.
Despite the investment and promise of plant-associated microbiome research, typical approaches are focused on examination of individual isolates or field studies of complex native communities. There exists a need for a reproducible, controllable and observable lab scale model that accurately represents field conditions. Our solution to this is the EcoFAB with a focus on four key objectives:
- Observability: to measure biogeochemical processes with unprecedented resolution on nested scales. This would enable a theoretical modeling science known as mesoscale systems biology.
- Reproducibility: to recapitulate the system and its dynamics at will. Because this is a biological system, there will be variation, but this variation must be manageable as it is in model organism developmental biology. A reproducible system is required for testing predictions and evaluating models.
- Controllability: to make defined, targeted perturbations or manipulations to the cohort of species, their spatiotemporal community structures or dynamics, and their genetic complements – all in biochemical and geophysical context. The ability to manipulate our system is a prerequisite for hypothesis testing.
- Ecosimilarity: to assess the extent to which our bench-top model reproduces key behavior observed in open-field, real-world systems. Ultimately, the lessons we learn must be transferable to extant ecologies to enable interventions promoting ecoremediation and a healthier planet.